What is PCR (polymerase chain reaction)?
Image credit: Greg Moss / Wellcome Sanger Institute
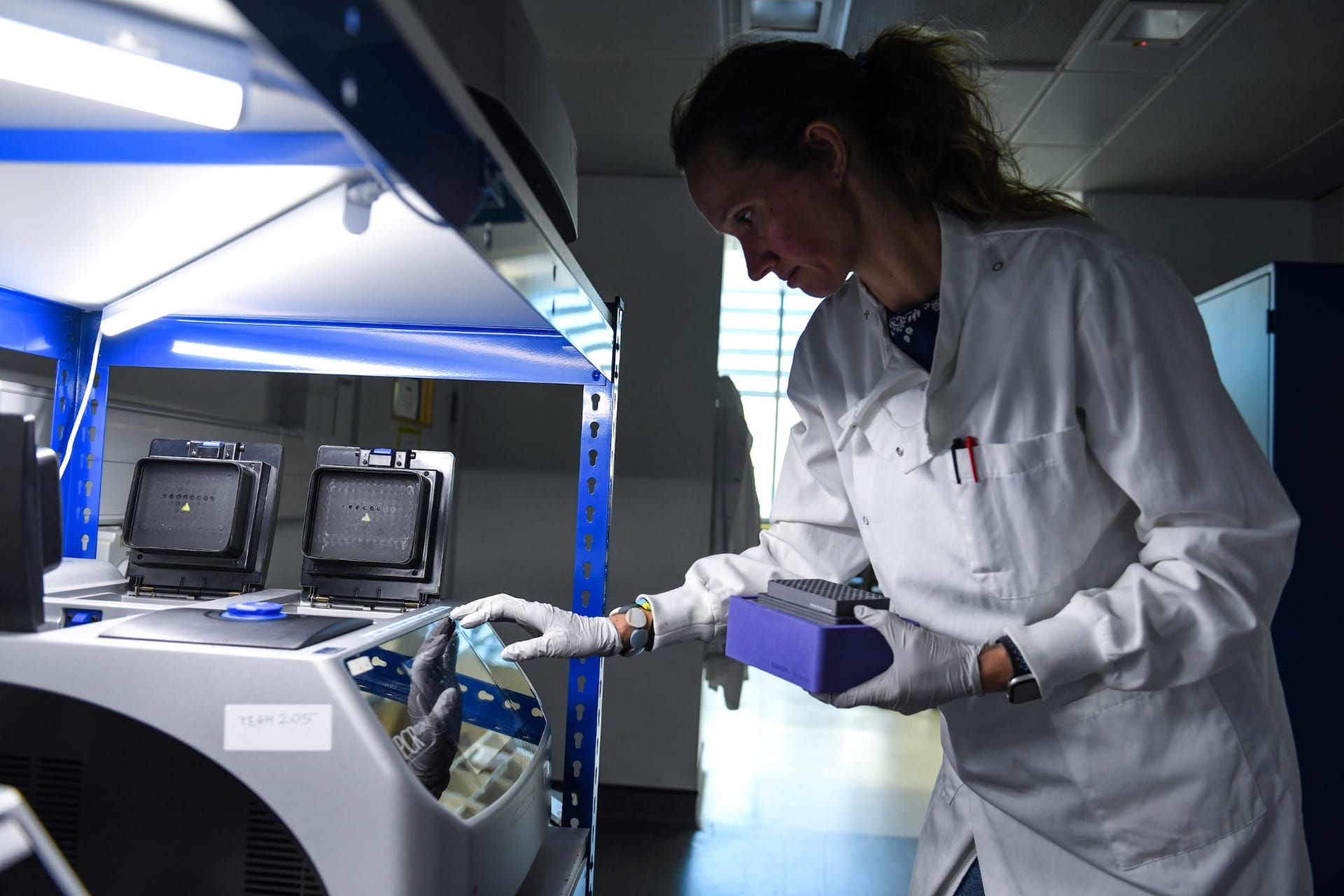

PCR is a technique used in the lab to make millions of copies of a particular section of DNA.
- PCR is used in molecular biology to make many copies of (amplify) small sections of DNA or a gene.
- This makes it possible to generate millions of copies of a particular section of DNA from a very small original amount.
- The polymerase chain reaction (PCR) was originally developed in 1983 by the American biochemist Kary Mullis. He was awarded the Nobel Prize in Chemistry in 1993 for his pioneering work.
- PCR was used as part of testing efforts during the Covid-19 pandemic to confirm whether a person was infected with the virus.
What is PCR (polymerase chain reaction)?
- Polymerase chain reaction (PCR) is a technique used in medicine and molecular biology research to make many thousands or even millions of copies of a section of DNA, such as a specific gene.
- It has many uses, including the early stages of processing DNA for sequencing or when generating forensic DNA profiles from tiny amounts of DNA.
- It can also be used to detect the presence or absence of a gene to help identify pathogens during infection. For example, during the Covid-19 pandemic, PCR tests were used to detect whether a person’s swab sample had a segment of the virus’s genetic material, to provide a positive or negative infection result.
How does PCR work?
Whatever the sample or application – the principles behind every PCR are the same.
A PCR has five core ‘ingredients’:
- the DNA template to be copied.
- primers – short stretches of DNA or RNA, 20 to 30 bases in length, that bind either side of the DNA section of interest and mark the point that PCR starts.
- DNA nucleotide bases (also known as dNTPs). DNA bases (A, T, C and G) are the building blocks of DNA and are needed to construct the new strand of DNA.
- an enzyme, called Taq polymerase, which adds the bases to the copied sequence.
- a buffer to ensure the right conditions for the reaction.
PCR has three main stages, which involve a process of heating and cooling:
- Step 1, denaturing: the double-stranded template DNA is heated, which separates it into two single strands.
- Step 2, annealing: the temperature is lowered to enable the DNA primers to attach to the template DNA.
- Step 3, extending: the temperature is raised again and the new strand of DNA is made by the Taq polymerase enzyme.
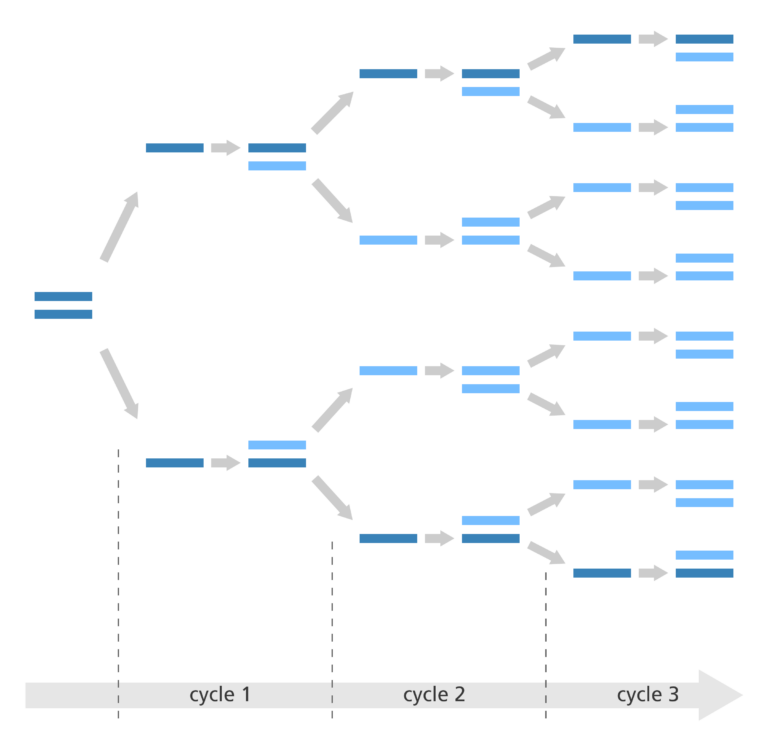
- These three stages are repeated 20-40 times, doubling the number of DNA copies each time. This is called thermal cycling, carried out by a machine.
- It’s carried out by a machine called a thermal cycler. Depending on the speed of the machine, PCR can be performed in under and hour or up to a few hours.
- After PCR has been completed, a method called electrophoresis can be used to check the quantity and size of the DNA fragments produced.
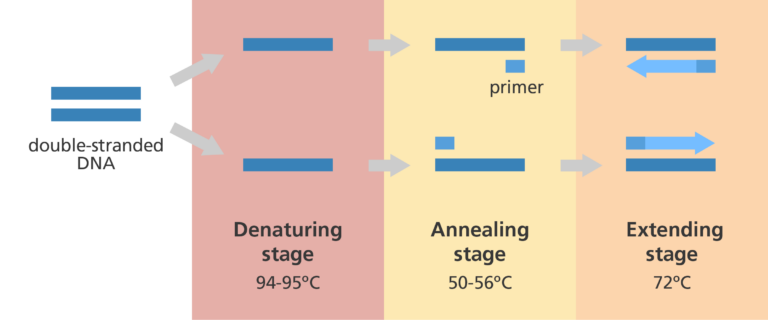
What happens at each stage of PCR?
Step 1: Denaturing
- The reaction mixture is heated to 94-95⁰C, for between 15 and 30 seconds.
- The high temperature causes the hydrogen bonds between the bases in two strands of template DNA to break and the two strands to separate.
- This results in two single strands of DNA, which will act as templates to produce the new copies of each strand of DNA.
- It is important that the temperature is maintained at this stage for long enough to ensure that the DNA strands have separated completely.
Step 2: Annealing
- The reaction is cooled to enable the primers to attach to a specific location on the single-stranded template DNA by way of hydrogen bonding.
- The temperature depends on the characteristics of the primer, but is usually between 50 and 65⁰C.
- The two separated strands of DNA are complementary and run in opposite directions (from one end – the 5’ end – to the other – the 3’ end). As a result, there are two primers – a forward primer and a reverse primer.
- This step is essential because the primers serve as the starting point for DNA synthesis, by providing a short region of double stranded DNA for the polymerase enzyme to work with. Only once the primer has bound can the polymerase enzyme attach and start making the new complementary strand of DNA from the loose DNA bases, in the extending step.
- The annealing step usually takes about 10-30 seconds.
Step 3: Extending
- The heat is increased to 72⁰C to enable the new DNA to be made by a special Taq DNA polymerase enzyme which adds DNA bases.
- Taq DNA polymerase is an enzyme taken from the bacteria Thermus aquaticus (“Taq”):
- This bacterium normally lives in hot springs so can tolerate temperatures above 80⁰C, but its optimum temperature is 72⁰C.
- The bacteria’s DNA polymerase is very stable at high temperatures, which means it can withstand the temperatures needed to break the strands of DNA apart in the denaturing stage of PCR.
- DNA polymerase from most other organisms would not be able to withstand these high temperatures. For example, human polymerase works ideally at 37˚C (body temperature).
- At 72⁰C, the Taq polymerase begins to build the complementary strand. It attaches to the primer and then adds DNA bases to the single strand one-by-one in the 5’ to 3’ direction.
- The result is a brand-new strand of DNA and a double-stranded molecule of DNA.
- The duration of this step depends on the length of DNA sequence being amplified. It usually takes around one minute to copy 1,000 DNA bases.
Repeating the process
- These three processes of thermal cycling are repeated 20-40 times to produce lots of copies of the DNA sequence of interest.
- The new fragments of DNA that are made during PCR also serve as templates to which the DNA polymerase enzyme can attach and start making DNA.
- The result is a huge number of copies of the specific DNA segment produced in a relatively short period of time.