Malaria: an ongoing battle against drug-resistance
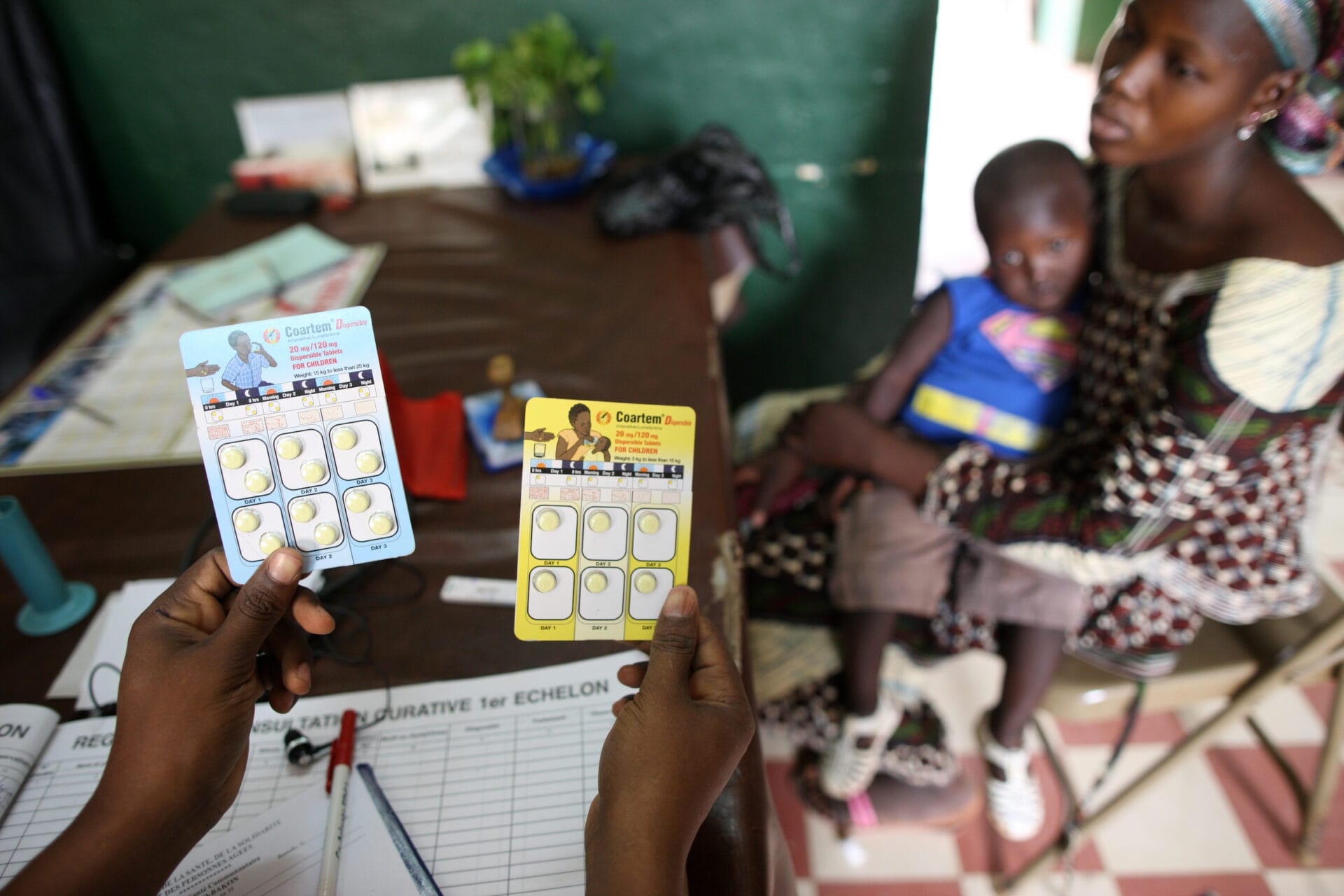

Resistance to antimalarial drugs is one of the biggest problems facing malaria control. Recent studies looking at the genome of the malaria parasite are helping scientists understand how drug resistance evolves – and how we could keep it in check.
This is part 2 in our series on ‘Using Genomics to Tackle Malaria’ series. Read part 1 here.
- Thanks to renewed efforts to prevent and control malaria, deaths from the disease have fallen by 47% since 2000.
- However, malaria is still widespread in more than 90 countries and kills around 600,000 people around the world each year. Controlling it remains a global priority.
- Scientists are using genomics to understand how the parasite that causes malaria can become resistant to treatment, so we can stay one step ahead.
Key terms
Genome
The complete set of genetic instructions required to build and maintain an organism.
Evolution
The process by which organisms change and adapt by natural selection.
Mutation
A change in the DNA sequence, which may have positive, negative or neutral effects on the organism.
Drug resistance in malaria parasites
Since 2000, deaths resulting from malaria have fallen globally by 47%. This is thanks, in part, to a boost in funding for malaria control – including raising awareness of prevention strategies, distributing mosquito nets and spraying insecticides to kill mosquitoes.
Antimalarial drugs have also played a major role in helping to control the disease, by quickly clearing the parasites that cause malaria from a person’s body. However, an ongoing challenge remains that the parasites continue to evolve and become resistant to these drugs – making the drugs less effective at treating the disease.
In fact, Plasmodium falciparum – the most serious group of Plasmodium species that can cause malaria – has developed resistance to nearly all antimalarial drugs currently in use.
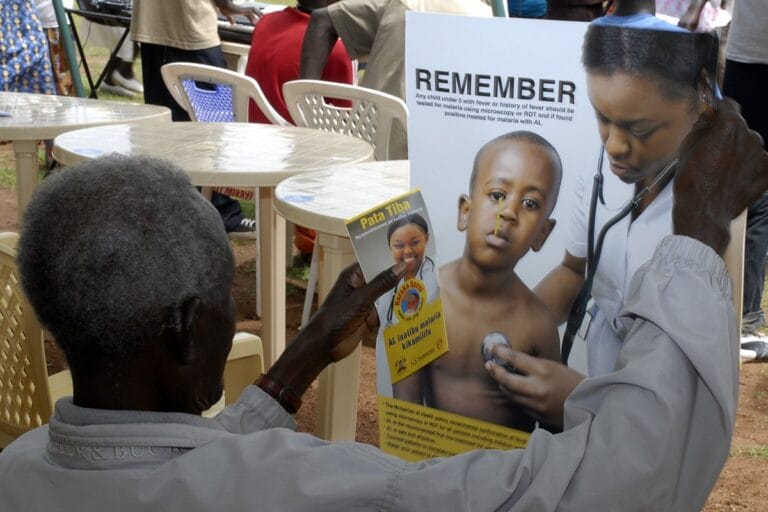
An evolutionary arms race: drug resistance in malaria parasites
To treat malaria and prevent recurrence, every single Plasmodium parasite needs to be removed from the body.
But it can take just one mutation to make a Plasmodium parasite better at avoiding the effects of a drug. Over time, when exposed to a drug, these parasites outcompete their neighbours and eventually become the dominant type in the population.
Once back inside the mosquito – the ‘vehicle’, or vector, through which Plasmodium spreads malaria between people – the parasite can pass their drug-resistant characteristics to the next generation, which enters a person’s blood when the mosquito bites. This can make antimalarial drugs much less effective at clearing the parasite from a person’s body – either acting too slowly or having no effect.
A hotspot of antimalarial resistance
Interestingly, resistance often emerges in the Greater Mekong Subregion in South East Asia, including in Cambodia, Laos, Myanmar, Thailand and Vietnam.
This might be because malaria infections are less common here than other places, meaning drug-resistant Plasmodium faces less competition from other parasites. Alternatively, people might develop natural immunity in areas where malaria is more common, like sub-Saharan Africa. This translates to a lower proportion of people being treated with antimalarial drugs, and so puts less drug pressure on the parasite.
Localised drug resistance is detrimental on a regional scale – but it can be devastating if the resistant parasite spreads to other parts of the world. This happened in the 1980s and 1990s, when a Plasmodium strain that was resistant to the drug chloroquine emerged in the Greater Mekong Subregion, before spreading to India, then to East Africa, and eventually all of Africa. Millions of people with this chloroquine-resistant malaria no longer responded to available treatment options, causing a dramatic increase in the number of malaria-related deaths.

Antimalarial drug resistance often emerges in the Greater Mekong Subregion in South East Asia, including in Cambodia, Laos, Myanmar, Thailand and Vietnam.
The global research community has battled drug resistant malaria for over a century:
1
Quinine
1632: the first recorded use of quinine for malaria
1910: the first recorded quinine-resistant Plasmodium
2
Chloroquine
1945: chloroquine is introduced as an antimalarial drug
1957: the first recorded chloroquine-resistant Plasmodium:
3
Sulfadoxine
1967: sulfadoxine pyrimethamine is introduced as an antimalarial drug
1967: the first recorded sulfadoxine pyrimethamine-resistant Plasmodium
4
Mefloquine
1977: mefloquine is introduced as an antimalarial drug
1982: the first recorded mefloquine-resistant Plasmodium
5
Artemisinin
1970s: artemisinin is introduced as an antimalarial drug
2009: the first recorded artemisinin-resistant Plasmodium
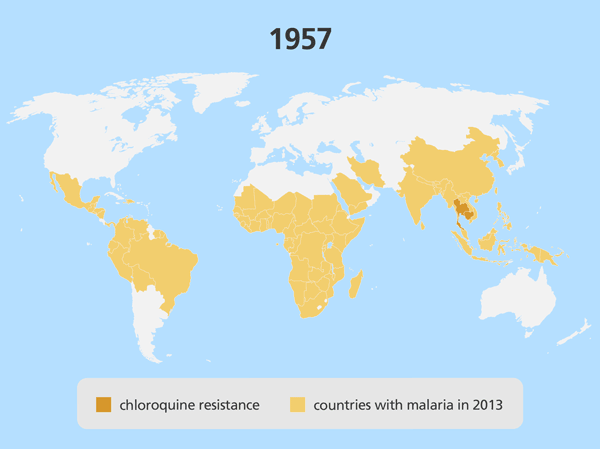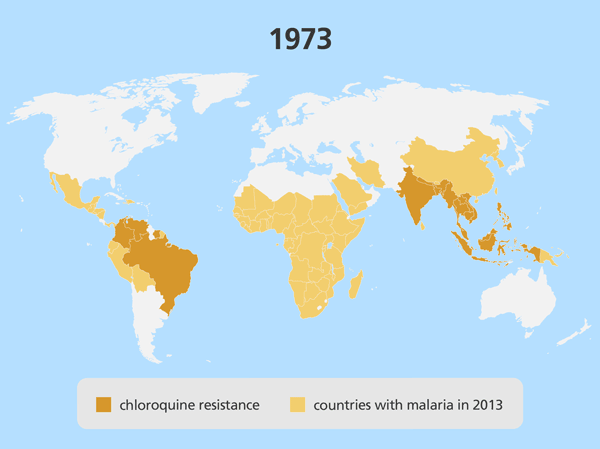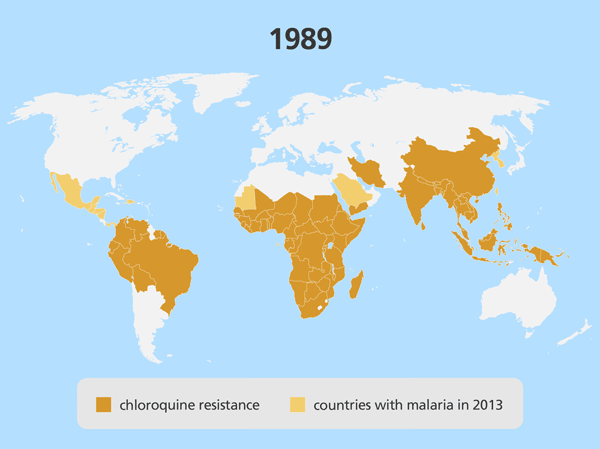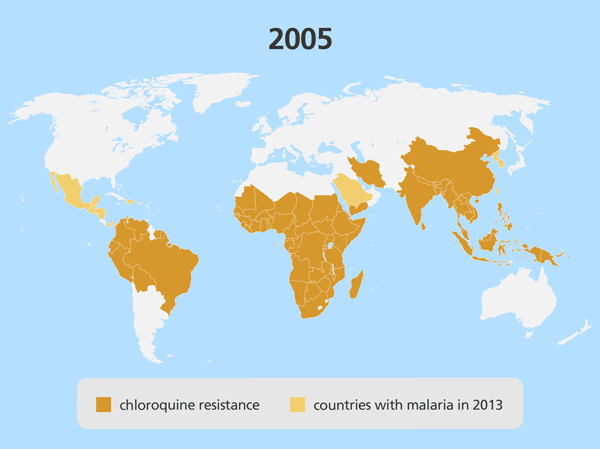
Combining treatments to overcome resistance to artemisinin
In 2009, researchers in the Greater Mekong Subregion noticed a worrying sign of resistance to artemisinin – a powerful antimalarial treatment that can normally reduce the number of Plasmodium parasites in the blood by half within just 12 hours.
Scientists knew that if the resistant parasite spread beyond the region, or emerged independently elsewhere, the public health consequences could be catastrophic – potentially reversing the decline in death from malaria that we’ve seen in recent decades.
In response, the World Health Organization launched emergency action, combining artemisinin with other drugs, known as artemisinin-based combination therapy (ACT). The fast-acting artemisinin first reduces the number of parasites, before the second partner drug eliminates any that are left.
The combination strategy proved useful at keeping the resistance contained to the region, particularly when the use of different ACTs was rotated. As of 2022, the World Health Organization recommended six different ACTs as treatment for people with different types of Plasmodium infection.

The World Health Organization’s goal is to remove malaria completely from Greater Mekong Subregion countries by 2030.
Genomics vs resistance
Thanks to advances in genome sequencing techniques, we know much more about the underlying genetic changes that have caused some strains of malaria parasite to become resistant to artemisinin. Often, the answer lies with the kelch13 gene, found on Plasmodium falciparum’s chromosome 13.
Scientists discovered this through several genetic experiments, including genome-wide association studies that compared thousands of Plasmodium falciparum genomes from across malaria hotspots in Africa and South East Asia. Another team grew the parasite in the lab for five years, occasionally exposing the colony to artemisinin to gradually encourage the development of resistance.
In all cases, they found that artemisinin resistance wasn’t caused by just one mutation in kelch13 – it was caused by more than 20.
Kelch13 and friends
The kelch13 gene is one of the most conserved genes in the Plasmodium genome. Before the use of artemisinin, kelch13 had remained pretty much unchanged for more than 50 million years, suggesting it’s crucial for the parasite’s survival – so it was surprising to find it with so many mutations.
The scientists wondered if a mutation in such an important gene might be detrimental to the parasite’s overall health. For example, does it make the parasite slower, less agile and less able to compete with its neighbours when the drug isn’t around?
Investigating further, they discovered mutations in four other genes that nearly always happened when kelch13 was mutated: fd, arps10, mdr2 and crt. It’s likely that these are background mutations that don’t directly cause drug resistance – but instead compensate for a fault in kelch13 to keep the parasite healthy.
This could inform important new intervention strategies in the future, should the parasites become resistant to the combination of drugs used in ACTs. For example, in 2022, several research studies and reports suggested that looking for this fixed set of mutations in Plasmodium samples from people with malaria could act as a warning sign that the parasites are at high-risk of developing artemisinin resistance.
Information like this could enable healthcare workers to target these areas with medicines or insecticides to kill any artemisinin-resistant parasites before they take hold and spread any further.
Genomics: a powerful tool in controlling infectious disease
Genomics is powerful. Not only can we use it to understand how pathogens evolve to become resistant to treatments, but we can also use this information to plan our next move. Like a spy, it can give us the intelligence we need to track emerging resistance and plan a counterattack before it becomes more widespread.
